Category: Parkinson's Disease: Pathophysiology
Objective: To investigate the correlations between serum metabolites with fecal microbiome, fecal metabolites, and clinical phenotypes in patients with Parkinson’s disease (PD).
Background: There is accumulating evidence showing altered gut microbial composition and metabolism in PD patients [1,2,3,4]. However, the impact of these gut changes on host metabolism, and vice versa, remains unclear.
Method: 67 PD patients and 64 non-PD controls were extensively phenotyped. Fecal and serum metabolomics were performed using liquid chromatography-mass spectrometry, while fecal microbiome was analysed using 16S rRNA gene sequencing. Multivariate analyses with false discovery rate (FDR) corrections were used to analyze between-group differences and correlations in this clinico-biological meta-dataset.
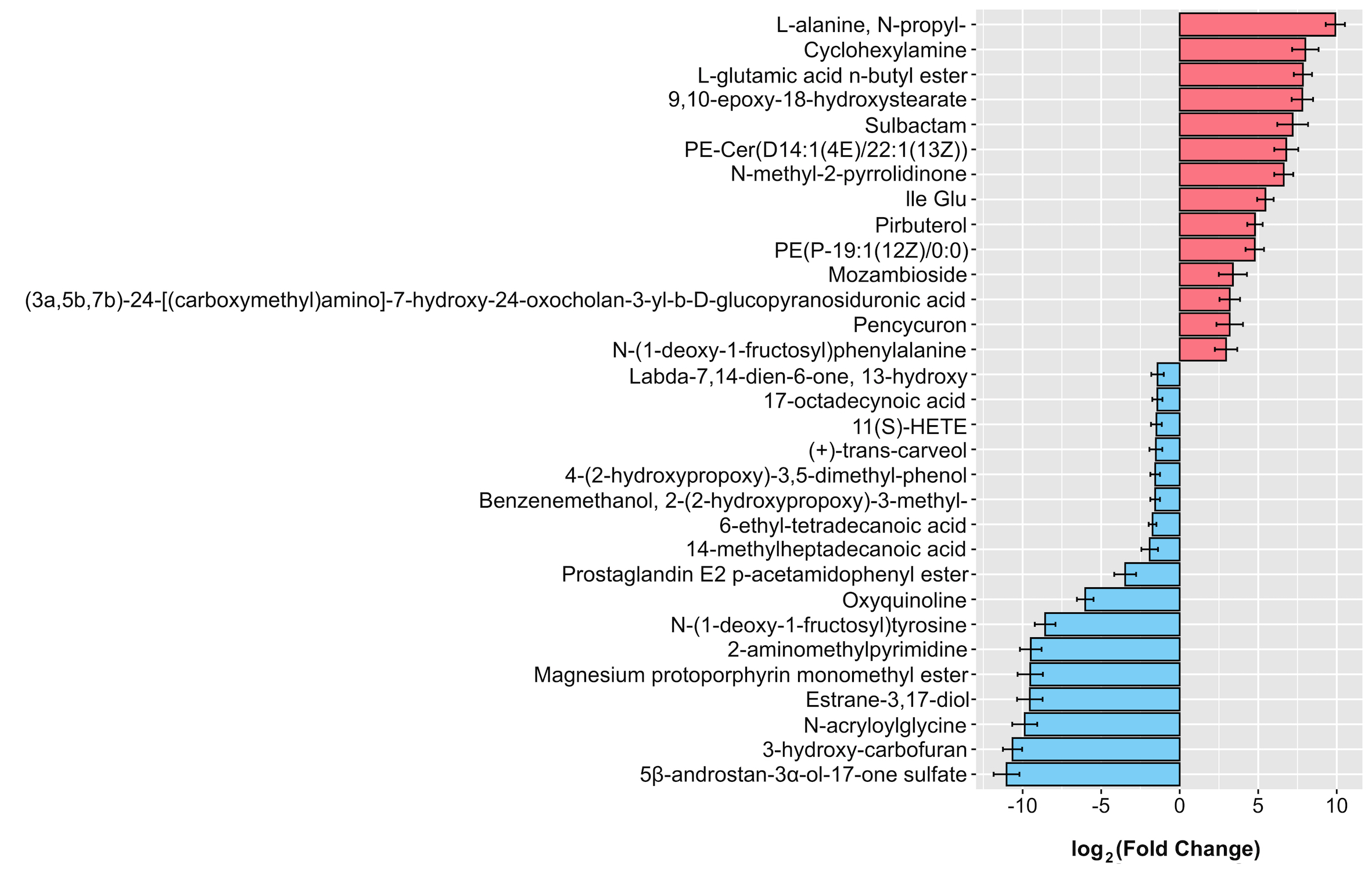
Results: Of the 473 filtered serum metabolites, we found 46 differentially abundant metabolites in patients vs. controls. These include alterations in lipids (especially fatty acyls, sterol lipids and glycerophospholipids), amino acids and organic compounds (including xenobiotics from pesticides, chemical solvents, drugs, food constituents, and natural products) [figure1]. Predictive functional analyses revealed consistent results — 23 pathways were altered in PD, with most of them involving metabolism of amino acids, lipids, hormones, vitamins and xenobiotics. Multivariate correlation analyses in the PD subgroup revealed associations between several serum and fecal fatty acids and phospholipids, as well as serum amino acid derivatives and specific gut microbes. Notably, the intercorrelated network of alanine, glutamic acid, pencycuron and phenylalanine in serum correlated with several fecal metabolites, microbial features and constipation severity in PD. Among the metabolites greatly reduced in PD (>9 log2 fold change), lower levels of 2-aminomethylpyrimidine correlated with worse Hoehn and Yahr staging, and is involved in the dipeptidyl peptidase-4 and glucagon-like peptide-1 pathway, believed to be neuroprotective in PD [5].
Conclusion: This multi-omics study revealed significant alterations in serum metabolome profiles of PD patients, and new insights concerning associations between host metabolites with gut metabolism, as well as with disease severity. Further studies are warranted to examine the role of these metabolites in PD pathophysiology.
References: [1] Tan AH, Lim SY, Lang AE. The microbiome-gut-brain axis in Parkinson disease – from basic research to the clinic. Nat Rev Neurol. 2022;18(8):476-495.
[2] Tan AH, Chong CW, Lim SY, Yap IKS, Teh CSJ, Loke MF, Song SL, Tan JY, Ang BH, Tan YQ, Kho MT, Bowman J, Mahadeva S, Yong HS, Lang AE. Gut microbial ecosystem in Parkinson disease: new clinicobiological insights from multi-omics. Ann Neurol. 2021;89(3):546-559.
[3] Wallen ZD, Demirkan A, Twa G, Cohen G, Dean MN, Standaert DG, Sampson TR, Payami H. Metagenomics of Parkinson’s disease implicates the gut microbiome in multiple disease mechanisms. Nat Commun. 2022;13(1):6958.
[4] Chen SJ, Chen CC, Liao HY, Wu YW, Liou JM, Wu MS, Kuo CH, Lin CH. Alteration of gut microbial metabolites in the systemic circulation of patients with Parkinson’s disease. J Parkinsons Dis. 2022;12(4):1219-1230.
[5] Bomba M, Granzotto A, Castelli V, Massetti N, Silvestri E, Canzoniero LMT, Cimini A, Sensi SL. Exenatide exerts cognitive effects by modulating the BDNF-TrkB neurotrophic axis in adult mice. Neurobiol Aging. 2018;64:33-43.
To cite this abstract in AMA style:
TS. Toh, CW. Chong, MF. Loke, JW. Hor, LC. Lit, J. Bowman, SL. Song, CSJ. Teh, S-Y. Lim, AH. Tan. Serum Metabolomics in Parkinson’s Disease: Correlations with Fecal Microbiome, Fecal Metabolomics and Clinical Status [abstract]. Mov Disord. 2023; 38 (suppl 1). https://www.mdsabstracts.org/abstract/serum-metabolomics-in-parkinsons-disease-correlations-with-fecal-microbiome-fecal-metabolomics-and-clinical-status/. Accessed May 6, 2026.« Back to 2023 International Congress
MDS Abstracts - https://www.mdsabstracts.org/abstract/serum-metabolomics-in-parkinsons-disease-correlations-with-fecal-microbiome-fecal-metabolomics-and-clinical-status/